Cauliflower Tales
Ron Petersen and Karen Hughes
Ecology & Evolutionary Biology, University of Tennessee
Knoxville, TN 37996-1100, USA
email: Ron Petersen, repete [at] utk.edu; Karen Hughes, khughes [at] utk.edu
From the Austro-Hungarian empire comes the saga of cauliflower mushrooms (Sparassis crispa and its allies) in tales of identity theft, sexual intrigue and revelation of most intimate secrets. Scenes shift to America, where the clan scatters across the continent, only to continue to produce new questions about immigration and political boundaries.
"Once upon a time…"
Like many a Jesuit, Father Franz Xavier Wulfen (1728-1805) mixed his theology with curiosity about the natural world. Having studied and taught mathematics in Vienna, he found himself in the town of Klagenfurt in what is today far eastern Austria. There he not only served his monastery, but investigated the local flora and fauna, but unlike many of the Brothers, he set about telling the world about his discoveries. His two-part report on the plants (for Wulfen and his peers, plants included fungi) was published by Baron N.J. Jacquin in Jacquin’s new journal revealing the newest scientific research in Austria. And thus we have the origin of “Clavaria crispa Wulfen ex Jacquin.” Wulfen’s color picture (Fig. 1) was fanciful at best, but even now it must be used as the basis for the name, longsince transferred as Sparassis crispa, the venerable European cauliflower mushroom.
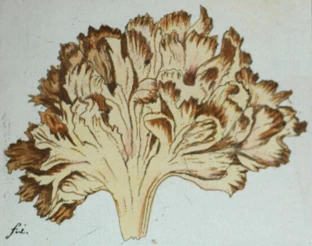
Fig. 1. Wulfen’s original lithograph of “Clavaria crispa.”
Original size 2 5/8 × 3 3/16 inches.
Truly, in greater Europe, no impersonator exists: S. crispa is unique to the eye, and Father Wulfen would be happy to know that his contribution is still honored however, he would not have suspected that in Europe, there were two interbreeding populations, perhaps the remnants of populations previously isolated by past glaciation.
But there’s more to the cauliflowers than meets the eye. Times change. One by one, new methods of inspection are applied — the hand lens (known in old literature as a “loupe”) gives way to the microscope, to which are added stains of various kinds, and most recently, an organism’s most intimate secret, its DNA, has come into use in identifying organisms of all kinds, including cauliflower mushrooms. Through it all, though, Wulfen’s name for S. crispa has persisted until now it is in use throughout the Northern Hemisphere from China and Russia, to North America and Europe.
International intrigue…
For some, “let sleeping dogs lie” might be applied to the cauliflower mushrooms. As long as we have a traditional name which can be applied widely, why not rest and go on to other more clear-cut problems? But the Jesuit in Father Wulfen would argue that if more can be understood, our curiosity should pursue. If new techniques are available, why not investigate what they can tell us? Even though cauliflower mushrooms exist widely in temperate forests, is there truly only one with such a widespread distribution? Or perhaps there is a patchwork of more limited entities which, with a little persistence, we can actually discern. Studies were based on herbarium and fresh collected materials. Spores for crossing studies were obtained by dropping spores from fresh material onto sterile aluminum foil, diluting and plating spores and culturing on agar for crosses.
Figure 2 shows what analysis of DNA sequences reveal about this heretofore monolith, Sparassis crispa. First of all, there are the remnants at least two major collection clusters under this name in Europe. Which of these populations was the original S. crispa? Look for IB75357 in the European cluster (arrow). That collection is from Klagenfurt and is the best we can do to place Father Wulfen’s original organism.
American collections fall into two well-supported clades, one from the Pacific Northwest and one broadly distributed clade from Eastern North America and Arizona. The diagram at the bottom of Fig. 2 shows our inferred genetic relationships within this latter group. There, the eastern sequences sort clearly into two sub-groups (eastern NA I and II). Surprisingly, however, a third cluster appeared between eastern NA I and II which represents first generation hybrids between the eastern NA I and II . There is no evidence of second generation (F2) hybrids and the first generation hybrids may be a dead end (see Cauliflower tale three). Just as interesting, a different branch (to the right) is populated only by sequences of collections from Arizona (more in Cauliflower tale two).
All this raises new questions. DNA analysis is not widely available, so can the major world-wide groups (not to mention the smaller subgroups) be separated by the naked eye or microscope? Just how different are the American entities (here named as S. radicata and S. americana and its form arizonica) from each other and from S. crispa? Meanwhile, Chinese researchers have successfully separated the most common Asian cauliflower mushroom and given it the name S. latifolia.
The most compelling reason for establishing the importance of DNA evidence in taxonomy is that while appearance (naked eye or microscope) is informative, DNA can provide additional evidence of evolution, biogeography and hybridization. So in spite of its general unavailability (and lingering mystery) for folks in the woods, DNA evidence is persuasive. But, confronted by clear-cut DNA evidence, could it be that mycologists have overlooked features which would have pointed to the same conclusion years before?
Cauliflower tale one…
Figure 3 is a photo of the cauliflower mushroom from the United Kingdom. In retrospect, it is important to notice the shape, size and color of the curly branch ends for comparison to American fruitbodies. The eastern North American organism can hardly be distinguished from the European in the field. Figure 4 is an eastern North American fruitbody from the Great Smoky Mountains National Park for comparison. Perhaps the curly branch tips are somewhat smaller and more compact than those of the European organism, but color differences are less than noticeable. What can’t be seen is whether the fruitbody has a significant, semi-gelatinous base which extends a few inches into the ground, not reported for the European entity.
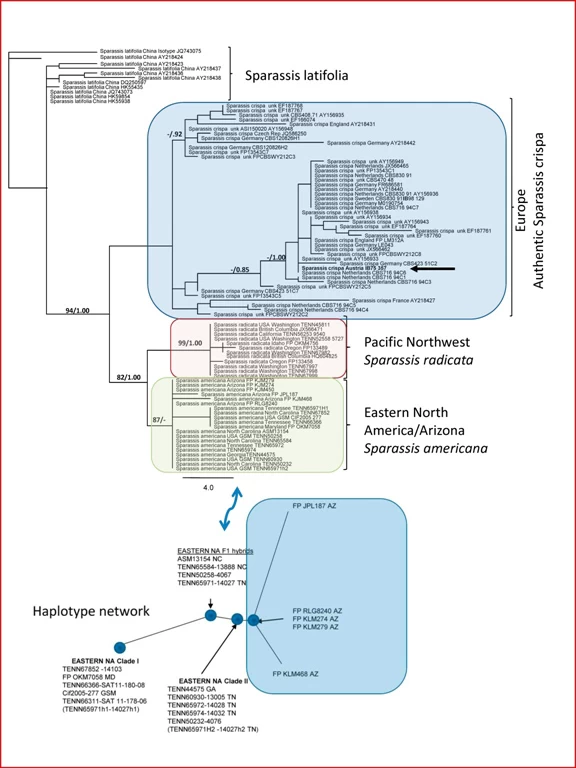
Fig. 2 Ribosomal ITS DNA phylogeny of the Sparassis crispa complex. One of 1000 most parsimonious trees of length 141. Ribosomal genes comprise, together with proteins comprise the ribosome; ITS is the internal spacer between ribosomal genes. This sequence has been selected as the fungal ‘barcode.’ Bootstrap support values greater than 70% are followed by Bayesian probability values greater than 0.70. Locations in the United States are given as state abbreviations. Unk = unknown location. GSMNP = Great Smoky Mountains National Park. C = clone number. Colored regions represent collections of the same species. The haplotype network minimizes base pair changes in the ribosomal ITS sequence between populations.
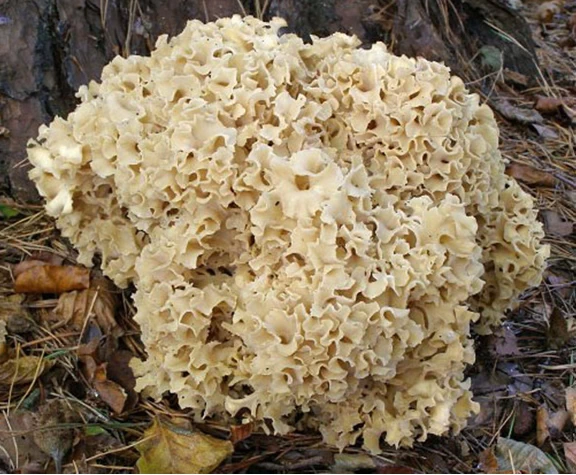

Fig. 3 Basidiome of Sparassis crispa from Europe, courtesy of Wikipedia.
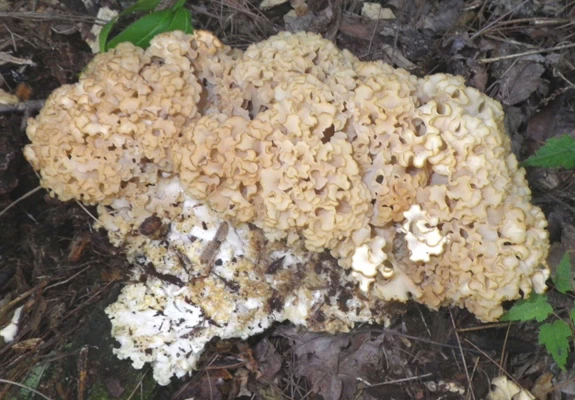

Fig. 4. Basidiome of Sparassis americana f. americana TENN65974 from Great Smoky Mountains National Park, TN. Photo by Dr. Rich Baird.

Fig. 5. Basidiome of Sparassis americana f. arizonica, courtesy of Flickr.
Back to the phylogeny in Fig.2. In the haplotype network, locate the large clade of American DNA sequences and notice that those of S. americana actually comprises two clusters, one of which seems purely from Arizona. Merely a glance at the photo (Fig. 5) reveals that the ultimate “curls” are small, finger-like frills — very fancy indeed — and that the fruitbody has a pinkish cast (not a photo artifact).
One doesn’t automatically think of Arizona as a classic habitat for fleshy fungi, much less cauliflowers, but in southern Arizona, east of Tucson, the tops of the “Sky Islands” of the Chiricahua Mountains are moist enough to support pine forests and fleshy fungi associated with them. A more careful look at the phylogeny shows that DNA sequences labeled as S. americana form arizonica really form a subset of the eastern cauliflower but apparently isolated in place long enough to form a group of its own.
Many years ago, a forest patholopgist named James R. Weir, working in the Pacific Northwest, noticed that whenever he came upon Sparassis, fruitbodies always sported a large, rooting stem. Accordingly, he named Sparassis radicata (Fig. 6). The name has suffered a clouded career, used by some, overlooked by others, but always available. A glance at the phylogeny in Fig. 2 shows that DNA clearly identifies the Pacific Northwest fungus as separate from S. americana. Indeed, the organism in the photo seems to have larger curls at the branch tips, and perhaps a slightly dull-yellowish cast, perhaps with age.
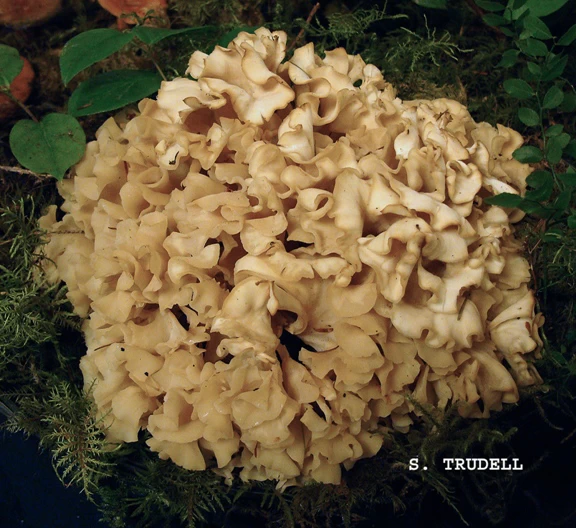

Fig. 6 Basidiome of Sparassis radicata, TENN67998-SAT12-295-03, courtesy Dr. Steve Trudell.
So it would appear that our first cauliflower tale ends with “everyone lives happily ever after” — the American Sparassis crispa entities can be recognized on sight (although S. americana is difficult to separate from S. crispa of Europe). Of course, in order to recognize them, it is necessary to be familiar with all of them, fresh in the field, and hopefully several times over. Travel and money, though, always create obstacles to such widespread exposure, and because all the Sparassises are edible and choice, many collections surely found their way into the cuisine rather than the herbarium.
Cauliflower tale two…
Not all of us are blessed to see all of these fungi fresh in the field, even with many years of collecting. The next best thing, albeit distinctly inferior, is a herbarium specimen — ideally accompanied by photos and thorough notes. Well documented herbarium specimens are surprisingly scarce (especially with such a group of edibles), but there were enough to tear apart for microscopic analysis. Can such autopsies support (or reject) what DNA tells us? If you’re not up for a little boredom, you might wish to skip to “Cauliflower tale three…” in which there is sexual intrigue.
Now, at the risk of too much terminology, a short anatomy lesson is in order. Those curly “petals” of the cauliflower mushroom are oriented with a lower side and upper side. Only the lower side produces basidia (the spore-producing structures; Figs. 7F, 8F, 10C, 11F) and basidiospores (Fig. 12 – but not to the same scale). Just above the basidia (going from lower surface upward) the hyphae of the flesh (Figs. 7A- C, 8A- C, 9A-F, 11A-D), of which there are several types. Just above the flesh of the petal there is a group of cells which are variously inflated and branched which are described as “free-form” (Figs. 7D, 8D, 9F, 10A, 11E). These free-form cells produce slender hyphae which cover the upper side of the petal like a lawn. Because they look like some structures called paraphyses, found in ascus-producing fungi, here they are called “paraphysoid hyphae” (Figs. 7E, 8E, 10B, 11F). All drawings were made by RHP.
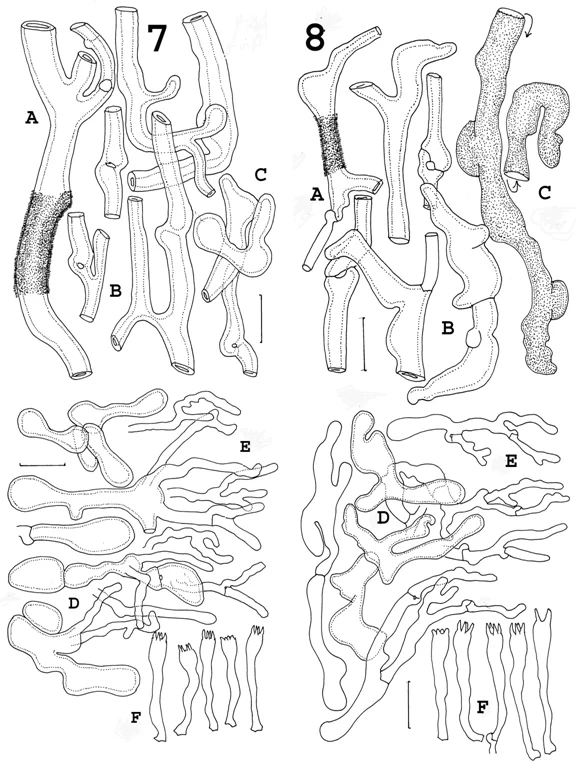
Fig. 7 Sparassis crispa. Tramal hyphae, origin of abhymenial paraphysoid hyphae and basidia. A. Rough-walled deep tramal hypha (only a portion shown as rough-walled). B. Deep tramal hyphae of middle branches, showing clamp connections. C. Articulation of deep and shallow tramal hyphae of upper branches. D. Inflated, thick-walled freeform cells of “dryophila structure.” E. Paraphysoid termini. F. Basidia. A-C, F 75/357 (IB). D, E. 1998/0129 (IB) Bars = 20 μm.
Fig. 8 Sparassis americana [f. americana]. Tramal hyphae of lower branches and stipe, origin of abhymenial paraphysoid hyphae, basidia. A. Rough-walled deep tramal hypha (only a portion shown as rough-walled). B. Deep tramal hyphae of middle branches, showing clamp connections. C. Thin-walled “vascular” hypha. D. Freeform hyphae of “dryophila structure.” E. Paraphysoid hyphae arising from subabhymenial cells. F. Basidia. A, B, E. TENN65974 (clade II). C. TENN50258 (clade II). D, F. TENN65584 (hybrid). Bars = 20 μm.
Figure 7 shows a montage of microscopic features of the European (true) Sparassis crispa. After all, it is the benchmark to which to compare American material. An early clue: keep your eye on two features, the “free-form” inflated cells which occur just under the surface of an individual branch (Fig. 7D) and those slender “paraphysoid hyphae” which cover the sterile upper surface of the branches (Fig. 7E). In European material (Fig. 7D), the “free-form” cells are often balloon-shaped — at least very wide; in fact, wider than those of any American specimens. When compared with those of S. americana (eastern form; Fig. 8D), differences are relatively obvious.
In S. americana f. arizonica (Fig. 9F), the “free-form” cells are more intricate and easily distinguished. The paraphysoid hyphae are shown in Fig. 10B. Fig. 11 shows the microstructures of S. radicata. The free-form cells (Fig. 11E), so prominent in other forms, are generally smaller and simpler in S. radicata, but the thick-walled tramal cells (Fig. 11C, D) include some inflated forms reminiscent of the European microstructures of true S. crispa in Fig. 7D. Altogether, not much to go on, and even then based on a lot of tedious hours at the microscope. The time versus profit ratio is somewhat dismal.
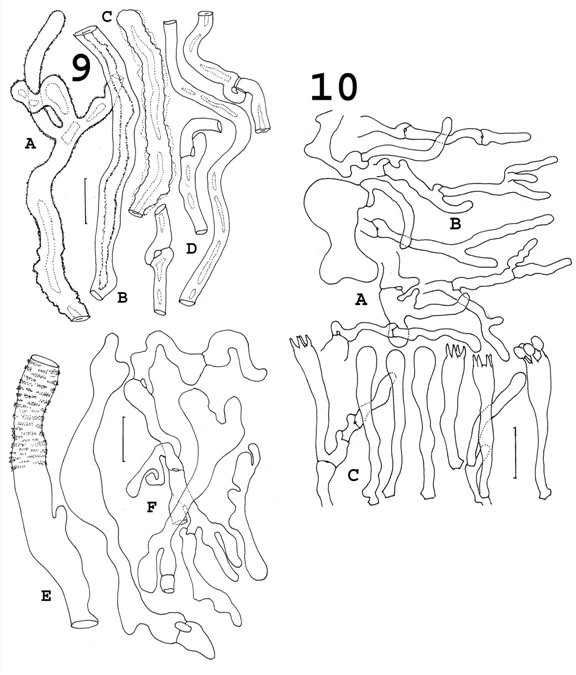
Fig. 9 Sparassis americana f. arizonica. Thick-walled, deep tramal hyphae, subabhymenial cells. a. Tramal hypha with outer wall roughened. b. Tramal hypha with smooth outer wall; inner wall roughened. c. Irregular gelatinous sheath surrounding tramal hypha; d. Tramal hyphae with smooth outer and inner walls, showing clamp connections. e. “Vascular hypha” of tramal tissue, showing banded ornamentation. f. Freeform subabhymenial cells. a-d. KJM 274 (CFMR). e, f. KJM 468 (CFMR). Bars= 20 μm.
Fig. 10 Sparassis americana f. arizonica. a. Paraphysoid hyphae arising from subabhymenial cells. b. Paraphysoid hyphae. c. Basidia and basidioles. a, c. KJM 468. (CFMR). b. KJM 274 (CFMR). Bar = 20 μm.
The other structures of note, remember, were the paraphysoid hyphae that cover the sterile surface of those curls at the ends of the branches. It takes a bit of doing, but three variations offer possibilities: the length of the structures (e.g. long in S. radicata, Fig. 11G), branching (e.g. S. crsipa, Fig. 7E) and internal septation (e.g. eastern S. americana, Fig. 8E).
Finally, often the most useful structures for comparison are spores. Fig. 12 shows the spores of all four organisms and proves that there isn’t a penny’s worth of difference in size and shape among them. Too bad, for this metric is usually a lot easier to harvest than those discussed above.
The end of the second Cauliflower tale might seem somewhat anticlimactic. While it is possible to distinguish specimens (fresh or dried) of the major groups in the phylogenetic tree microscopically, it may not be worth the effort. It may merely be a relatively inconvenient method to confirm the separation of the major clusters.
Cauliflower tale three…
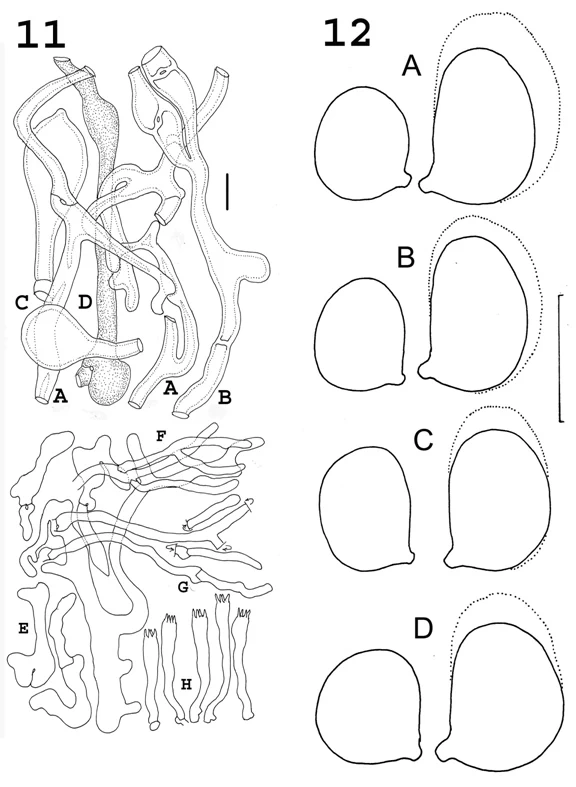
Fig. 11 Sparassis radicata. Microscopic structures. A, B. Thick-walled tramal hyphae. C Inflated thick-walled tramal hyphal segments. D. “Vascular” hypha. E. Freeform subabhymenial cells; F, G. Paraphyses. H. Basidia. A, C. TENN34723-RW004675. B, D. TENN67998-SAT12-295-02. E, G, H. TENN52558. f. TENN56253. Bar = 20 μm.
Fig. 12 Basidiospores of Sparassis crispa complex. A. S. crispa (Europe).B. S. americana [f. americana]. C. S. radicata. D. S. americana f. arizonica. Solid lines represent 90% of spore dimensions and shape. Stippled lines represent largest 10% of spore dimensions and shape. Bar = 5 μm.
One cliché tells a lot: Sex abounds.
Of the ways to define that illusive word “species,” one championed by Harvard Professor Ernst Mayr can be abbreviated to “If they can successfully interbreed, they belong to the same species.” The implication was that appearance (naked eye or microscope) was distinctly subordinate to mating behavior and production of offspring. We can quibble about “successfully,” but in the Sparassis crispa complex, here’s our application of the principle. If two haploid isolates (read sperm and egg if need be) can join together and the resulting fungal hyphae form those little structures called clamp connections (they are a proxy for sexual compatibility in many higher fungi), we consider the parents to potentially belong to one sexual species. However, the ability to mate alone is insufficient. Horses and donkeys are different species but can mate. Lions and tigers likewise.
Armed with the DNA information in Fig. 2, will haploid strains of the members of the S. crispa complex recognize each other fondly (in spite of long-distance and long-time separation) and show us that they intend to interbreed, or will they reject one another, either universally or selectively? The saga of how to produce those haploid strains is for another day, but we were able to pair haploid strains as shown in Fig. 13. While sexual recognition and clamp production were not universal, every representative successfully paired with at least one other, but there are varying degrees of mating failure suggesting that reproductive isolation is in progress.
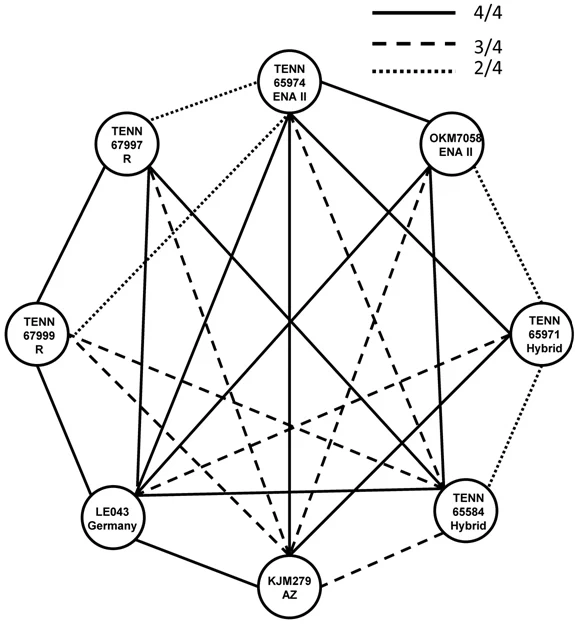
Fig. 13. Pairings between haploid (single spore) isolates of Sparassis crispa complex. Alphanumeric designations = representative strain. R = S. radicata; AZ = S. americana form arizonica; ENA II = clade two of S. americana. Hybrid = results of DNA analyses showing interbreeding among eastern North American S. americana. Pairings = four between each representative. Absence of a line indicates failure to produce clamp connections. Level of mating success is indicated by solid lines (4/4 matings were successful as indicated by the formation of clamp connections) to dotted lines (only 2/4 matings successfully formed clamps, an indication of partial reproductive isolation).
The take-home message from this brief excursion into sexual mores is that even though DNA shows unmistakable distinction among the members of the S. crispa complex, and even though the organisms can be recognized with the naked eye and under the microscope, the fungi themselves seem to overlook these differences and can potentially interbreed at some level when crossed in the lab. Whether they do so in nature or whether progeny are fertile or persist is uncertain. There is limited evidence that progeny may not persist in some cases (note for example the presence of F1 but not F2 progeny between eastern populations of S. crispa and lack of any detected progeny between Arizona and Eastern North American populations) If, in fact, were our assessment based solely on sexual compatibility in the lab, we would conclude that all of the S. crispa complex (we don’t know about S. latifolia) belong to one species but that would ignore what is essentially speciation in process.
The moral of these tales…
Aesop was always careful to conclude his fables with a moral. If you’re still conscious, perhaps the question “Who cares?” might have occurred. First, that old TV quiz show, “Who do you trust?” In the case of cauliflower mushrooms, do you trust appearances (naked eye or microscope), sexual compatibility, or the DNA evidence? Since appearance agrees with DNA that distinct taxonomic units exist between eastern North America and Europe, we feel comfortable naming S. americana as distinct — that is, NO LONGER correctly called S. crispa – as well as its form arizonica. Experience has shown us that sexual compatibility may be retained long after significant genetic differentiation between groups has occurred. As in all taxonomic decisions, however, it is a matter of judgment when significant differentiation implies speciation.
Moreover, the Jesuit in all of us can smile, realizing that now we know more about Sparassis crispa than we did before. There the story might end with “Angels and heads of pins,” but there is more. How many cauliflower mushrooms are there? We have discovered at least one (S. americana) heretofore unknown, which can be added to lists of species over the eastern United States. Sparassis radicata has been vindicated — it’s a good species.
Finally, the identification of cauliflower mushrooms is deceptively easy. Sparassis crispa seems limited to Europe, including western Russia, being replaced in eastern North America by S. americana and in western North America by S. americana f. arizonica in southeastern Arizona and S. radicata in the Pacific Northwest. The dominant species in Asia is S. latifolia. When you find a cauliflower mushroom, reckon where you are: each geographic population seems to have a separate, acceptable name. Use it.
And that’s the end of our story. Thanks for reading.
Acknowledgments
Most research is not done in isolation. Here are a few folks who helped. Dr. Steve Trudell and Mr. Brian Luther organized and furnished specimens and spore prints of S. radicata. Dr. Nadezhda Psurtseva furnished cultures from Komarov Botanical Institute, St. Petersburg, Russia. Ms. Rita Rentmeester expedited dikaryon cultures from Center for Forest Mycology (USDA Forest Service) in Madison, WI, which represented several collections cited by previous authors, and Dr. Beatriz Ortiz-Santana arranged a loan of specimens from CFMR, including material from the Sky Islands. Dr. Egon Horak investigated the herbarium in Innsbruck, Austria, and identified specimens of S. crispa from Klagenfurt. Numerous citizen scientists funneled specimens and spore prints to us individually or through others. Financially, research was supported, in part, by a National Science Foundation Grant to RHP and KWH.
This paper is based, in part, on a talk at NAMA 2013, and prior publication in Mycological Progress. You can catch it at: http://link.springer.com/article/10.1007%2Fs11557-013-0927-1#page-1
Citation: Petersen, Ron and Hughes, Karen (2014) Cauliflower Tales, McIlvainea 23 (13-15): https://namyco.org/cauliflower-tales/
Editor: Michael W. Beug
Publisher: North American Mycological Association
Published: May 1, 2014
Copyright: © 2014 Petersen, Ron and Hughes, Karen
NAMA Store >